This is part 1 of a 3 part series. Part 2 is here. Part 3 is here.
The human genome project completed the first draft of the human genome in 2000 after a 10 year push. Looking back, what’s striking is the expectation this draft genome would provide immediate practical use in medicine. President Clinton’s office sent out a press release at the time saying “Before the advent of the Human Genome Project….connecting a gene with a disease was a slow, arduous, painstaking, and frequently imprecise process. Today, genes are discovered and described within days.” The press release went on to gushingly list all the new medicine to come. Now some of this was merely hype to justify the $100 million research expense. But it also reflected a belief that the genetics of medical diseases would be straightforward. That turned out not to be the case.
The reality uncovered by the genome was most diseases are caused by large numbers of genes of very small effect. And these genes subtly interact with the environment in such a way that they only occasionally lead to disease. In retrospect, the simple view that single very bad genes were causing common diseases was based on a leap of faith that evolutionary selection doesn’t work very well. Bad genes cause genetic load, and in the case where a single gene by itself causes major disease the math says it should get weeded out efficiently by selection. All that said, in another 10-20 years with more computer power, more research, and massive amounts of genome data the medical promise should start to be fulfilled. It’s just a lot more work to isolate so many genes of small effect.
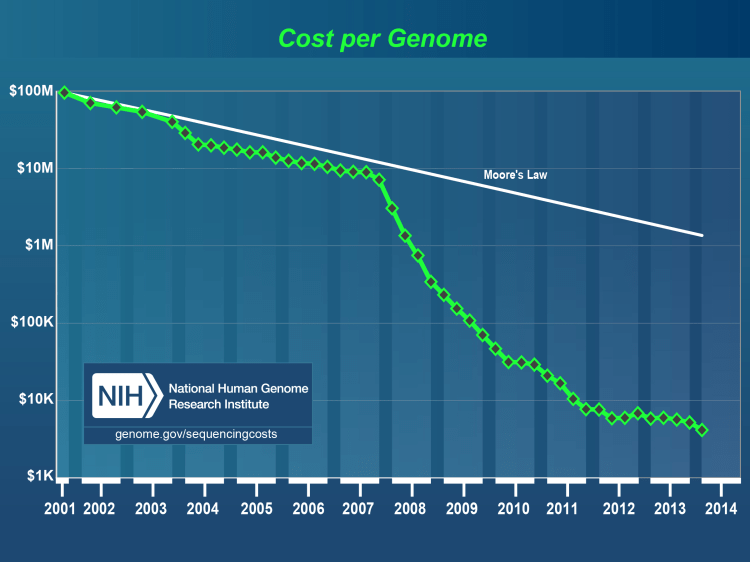
What’s amazing is how fast genome sequencing cost has fallen since 2000. I remember watching the movie Gattaca in the late 1990’s and thinking at the time it was a pretty good movie, but the writers didn’t know their science since it took $100 million and 10 years to type a genome. Not 30 seconds. But only a decade and a half later we’re well on our way there. This chart from the National Genome Research Institue tells the tale. The Moore’s Law line is a bit confusing until you realize it’s just there to baseline cutting costs in half every two years. Which is the trend in computer chips. So from 2001 to 2012 chip costs came down by a factor of 40. An incredible amount. Meanwhile genome sequencing costs came down by a staggering factor of 10,000. From $100 million per genome to $10k. Hundred dollar genomes will be here in another 5-10 years.
If medicine is not quite there yet, what is this technology good for? Well, the new cost structure opens up the possibility of genotyping all the major phyla of life on earth within our lifetimes. We’ll see how everything is related to everything else. It turns out that a bucket of ocean water has massive amounts of unknown bacteria, and by sequencing it we can start to understand how those bacteria are related. And the microbiome in your stomach has incredible unexpected diversity. We’re getting new details on how archaea evolved as the first forms of life.
What this data is especially useful for is evolutionary phylogeny. Determining how species are related to each other, and how they changed over time. If you know the DNA for many species, you don’t have to understand the gene functions themselves to figure out how the species are related. The math gets complex, but at a high level it’s reasonably straightforward. You compare DNA differences between species and those with the least change are the most related. You also have to know or estimate the underlying mutation rate, which gives an expectation of mutational change in DNA over time. Since most mutations at the molecular level are neutral, they have no impact on fitness and can be used as a molecular clock to show how long species have been separated. And with those two key ideas, we can figure out that whales are closely related to hippos, and the ancestral split occurred about 60 million years ago. We are living in a golden age for genomics biology.
Another good recent example is the relation between polar bears and brown bears. A new paper ”looked at the full genomes of polar bears, brown bears and black bears. Dr. Lindqvist, the senior author, said the full-genome approach enabled the scientists to look at polar bear history in new depth.” A study in 2010 set the species split at 600,000 years ago based on smaller snippets of DNA, under the evolutionary model of a single split. But the fuller analysis published just now showed the species split 5-6 million years ago, but have had some restricted gene flow between them ever since. This shows up in the data as recent rare occasional matches in the DNA between the species due to interbreeding (technically gene introgression), but with the majority of the DNA showing long term separation and neutral mutation. Which takes us to hominins.
The sequencing of ancient (40k years old) Neanderthal and Denisovan hominin DNA has surprisingly shown introgression into humans alive today. In fact the Denisovan DNA comes from a single finger bone, so once sequenced the fact it was a new hominin was startling. Regardless, what’s cool is the introgression pattern is similar to the polar bear case. Speciation followed by introgression. The old multiregional versus out of africa debate seems stale now that there’s real data to chew on. The newer model seems to be one where humans spread out and speciated across much of the globe, then further expansions led to waves of replacement with introgression from archaic forms. The 2000 year long Bantu expansion is turning out to be a good model. There’s similar work on the origin of Indo Europeans and the settlement of Europe showing the same pattern of replacement with introgression, but undoubtably the older school of thought saying most of it was cultural diffusion rather than replacement will win some of the day. What’s really shocking and new is that in principal archaic DNA can be used in conjunction with anthropology and archaeology to figure out how various waves of migration, replacement, cultural diffusion and introgression occurred in detail. Both in the past 10,000 years since the advent of agriculture as well as deeper into time when modern humans were replacing archaic forms.
As an aside, it’s worth mentioning that genetic markers can now be used to trace anyone’s ancestry at a personal level. 23andme is selling this now. To be fair the commercial reports aren’t that good yet. But they’ll get better as the 1000 genomes project and other similar efforts at deep global population sampling come on line. Genetically classifying the ancestry of all forms of life, including humans, is one of the hottest areas of science right now. This also means that Richard Lewontin was wrong in his 1972 paper on race and genetics, but in a very interesting way. In fact the future direction of the debate around Lewontin’s fallacy deserves a post of its own next week.
This is part 1 of a 3 part series. Part 2 is here. Part 3 is here.

3 comments